Methods and Applications of RNA Contact Prediction
Zhao group at Central China Normal University
Methods and Applications of RNA Contact Prediction
The RNA tertiary structure is essential to understanding the function and biological processes. Unfortunately, it is still challenging to determine the large RNA structure from direct experimentation or computational modeling. One promising approach is first to predict the tertiary contacts and then use the contacts as constraints to model the structure. The RNA structure modeling depends on the contact prediction accuracy. Although many contact prediction methods have been developed in the protein field, there are only several contact prediction methods in the RNA field at present. Here, we review the theoretical basis, advances, and limitations of some RNA contact prediction methods for tertiary structure and complex modeling problems. Moreover, we test the performances of these RNA contact prediction methods with two testing datasets.
Dataset-I:
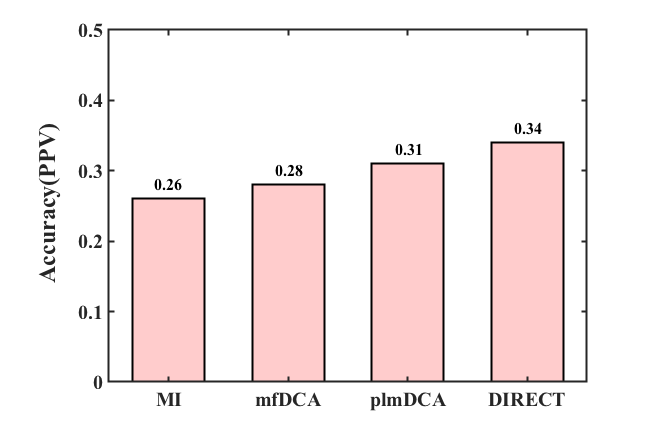
The dataset-I is a published riboswitch benchmark dataset for RNA intramolecular contact testing. As shown in Fig. 3, the results show that accuracy (positive predictive value, PPV) of MI, mfDCA, plmDCA, and DIRECT is 0.26, 0.28, 0.31, and 0.34, respectively.
Fig. 3. The accuracy of RNA intramolecular contact prediction methods. The accuracy (positive predictive value, PPV) of MI, mfDCA, plmDCA, and DIRECT are 0.26, 0.28, 0.31, and 0.34, respectively
Dataset-II:
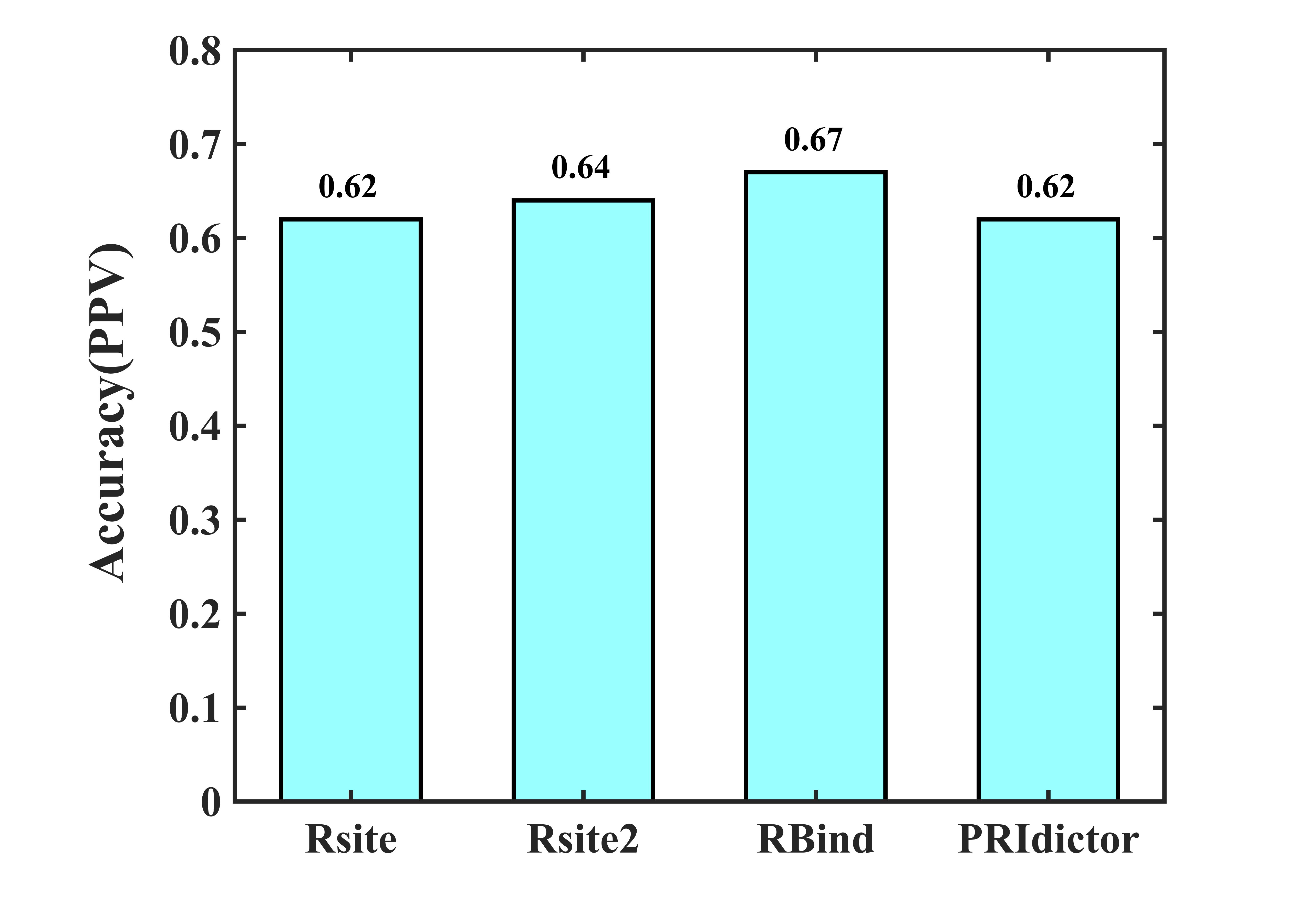
The dataset-II is an RNA-protein dataset (Table S1) for RNA intermolecular contact testing. The results show that the accuracy (positive predictive value, PPV) of Rsite, Rsite2, RBind, and PRIdictor is 0.62, 0.64, 0.67, and 0.62, respectively (Fig. 4).
Fig. 4. The accuracy of RNA intermolecular contact prediction methods. The accuracy (positive predictive value, PPV) of Rsite, Rsite2, RBind, and PRIdictor are 0.62, 0.64, 0.67, and 0.62, respectively.
Download
- The sequence and structure information of Dataset-I.
- The sequence and structure information of Dataset-II.
Copyright 2020, Lab of Biophysics