Statistics analysis results graph
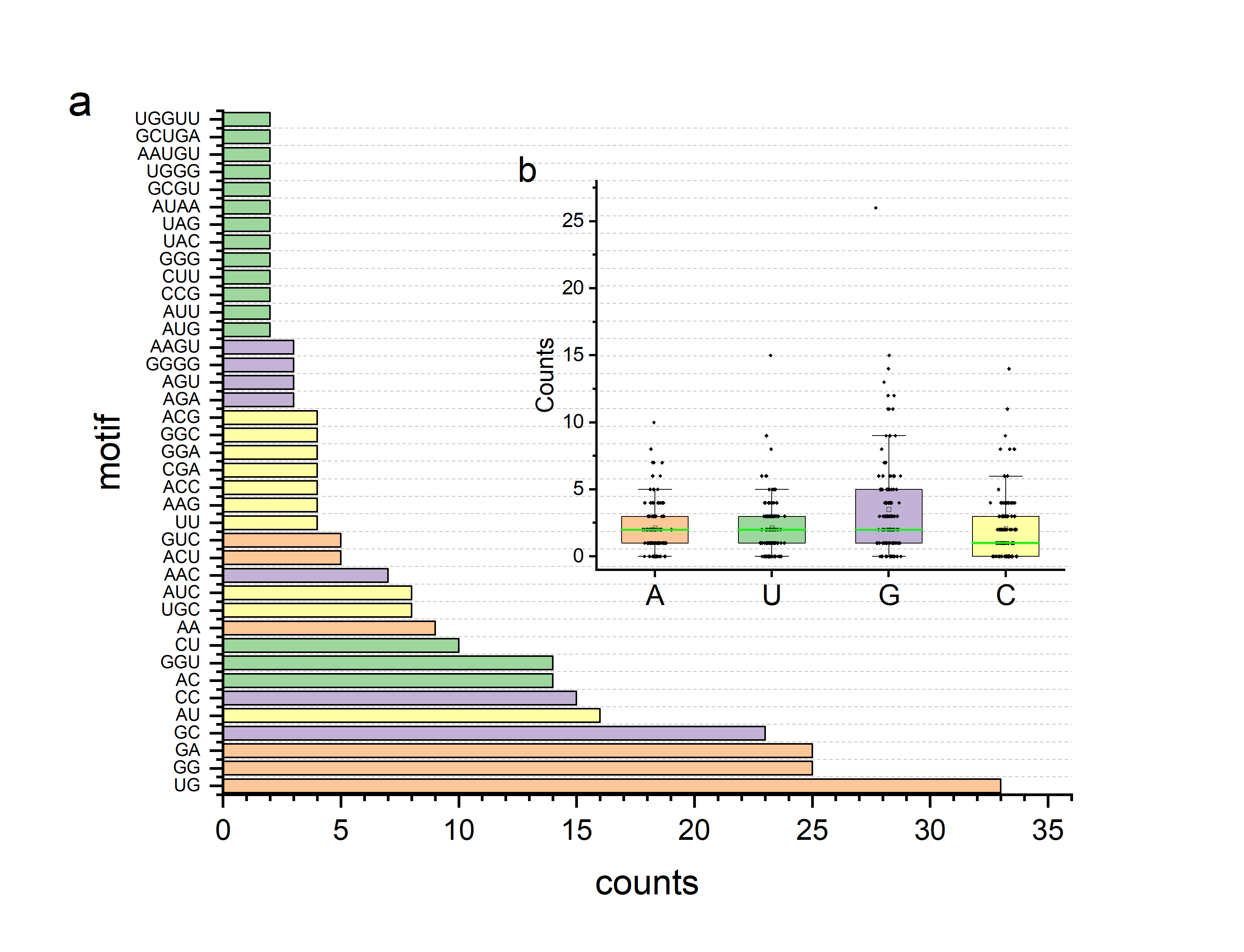
Figure 1. The motif and base distribution of binding sites of RNA-ligand. In Figure 1(a), the ordinate represents different motifs, and the abscissa represents the number of occurrences of the corresponding motif. In Figure 1(b), the abscissa represents four types of bases, and the ordinate represents the number of bases in each binding site.
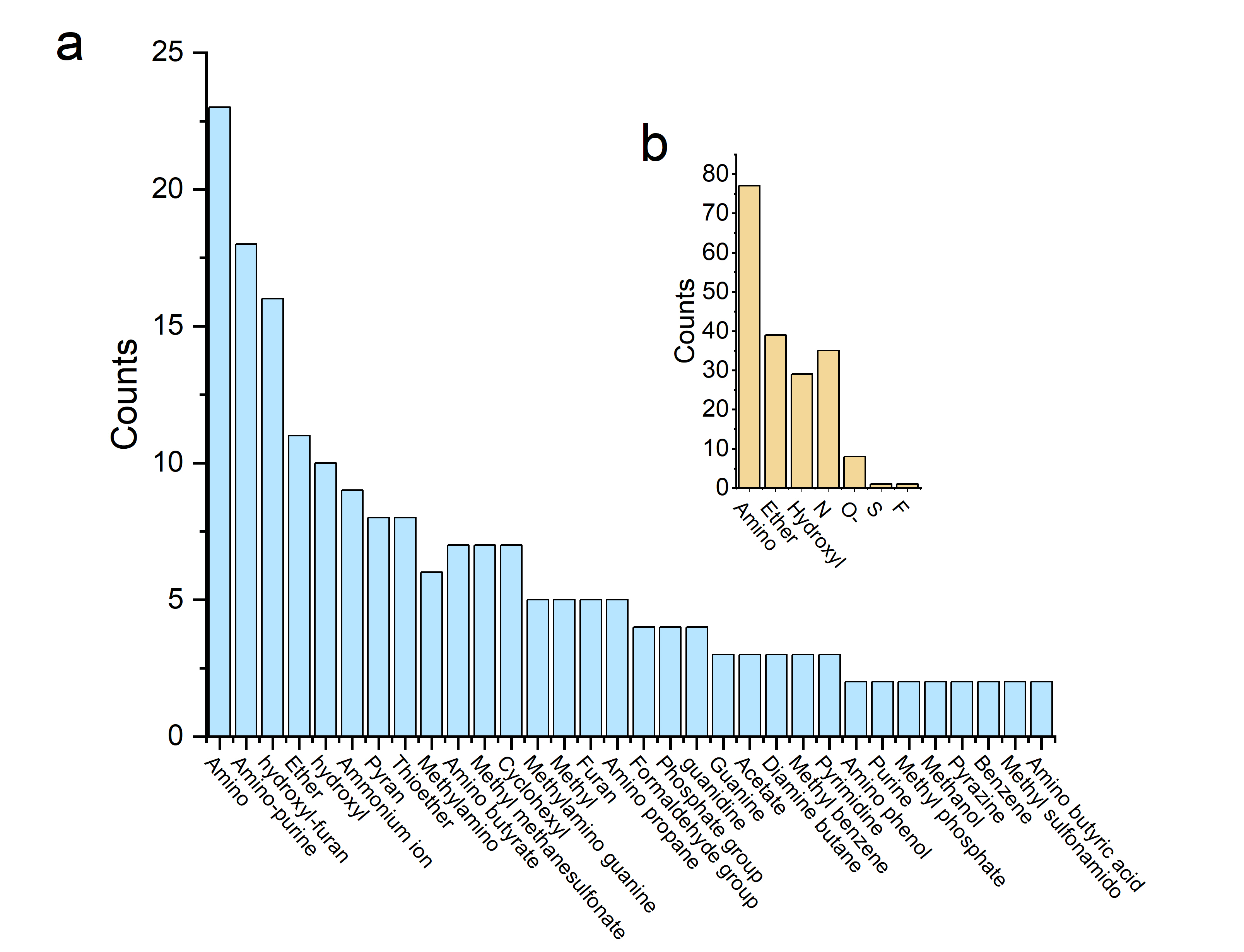
Figure 2. Hierarchical map of functional groups of ligands. Figure 2(a) is for hydrogen bond Interactions, and 2(b) is for non-bonded contacts. The ordinate represents the motifs of functional groups of ligands appeared in interactions, and the abscissa represents the number of the motifs.
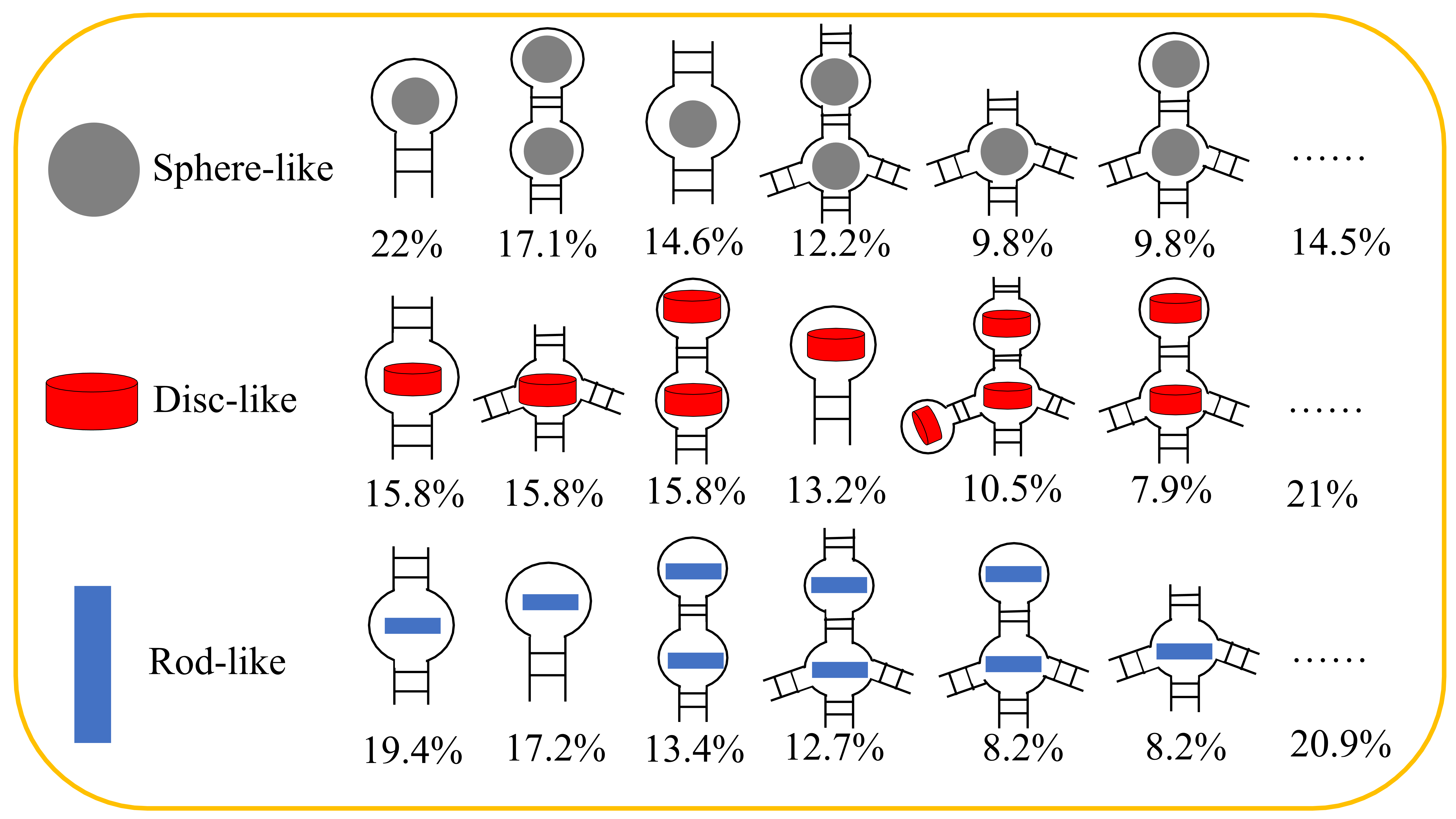
Figure 3. The binding patterns of the pockets on RNA secondary elements. The approximately spherical pockets are indicated by a gray circle, the disc-like pockets are indicated by a red pie, and the rod-like pockets are indicated by a blue rectangle. These representations are put into RNA topological diagrams further to show different binding patterns the percentage in all the binding patterns (bottom).
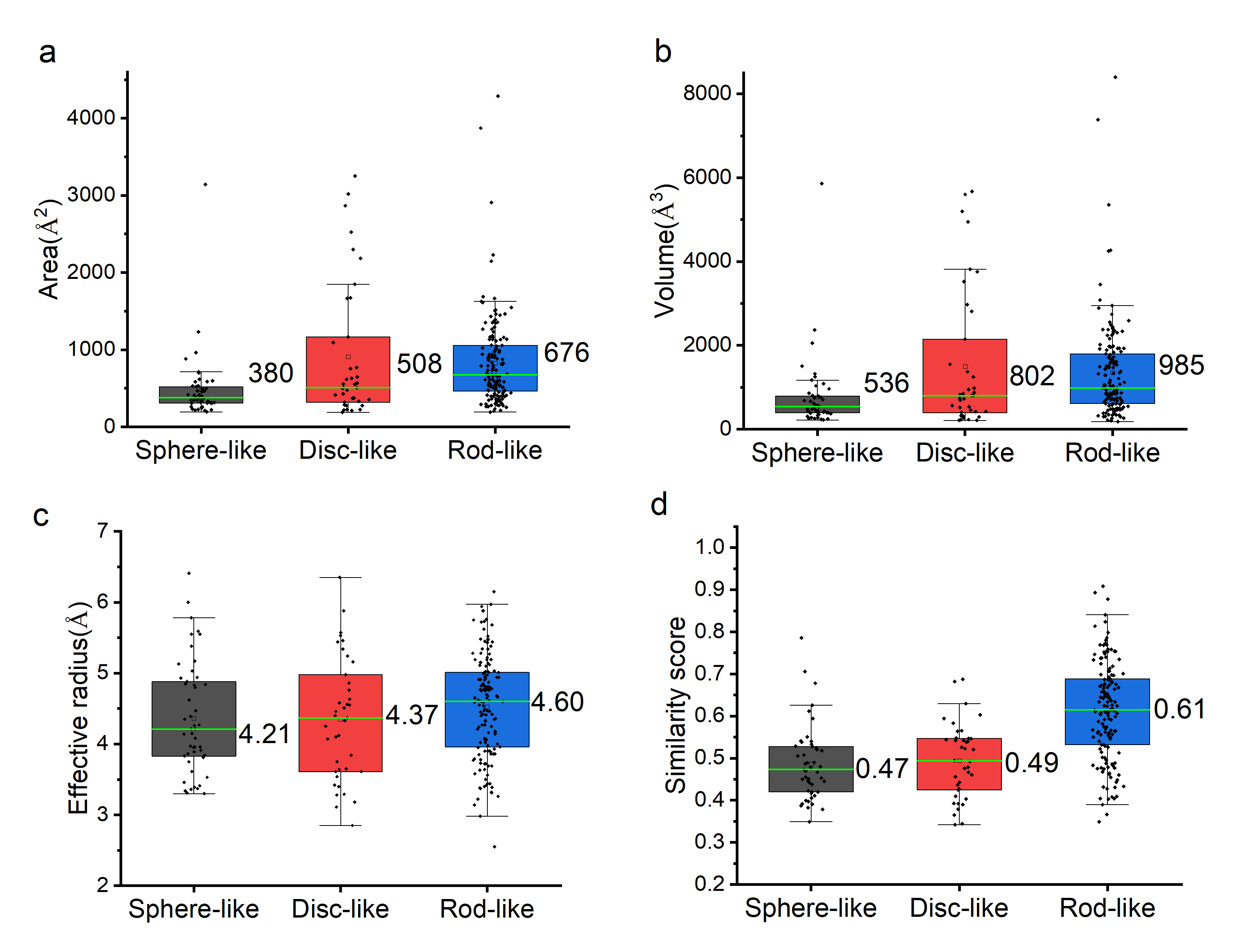
Figure 4. The geometric information distribution of the pockets among the three classes depicted by box and whisker charts (whiskers of boxes correspond to highest and lowest values, ends to upper and lower quartiles, and green line to median), Figure 3(a), 3(b), 3(c) and 3(d) depict the distributions of the area, volume, effective radius, and shape similarity scores of the three types of pockets correspondingly. Median values of each distribution are marked aside the box charts.
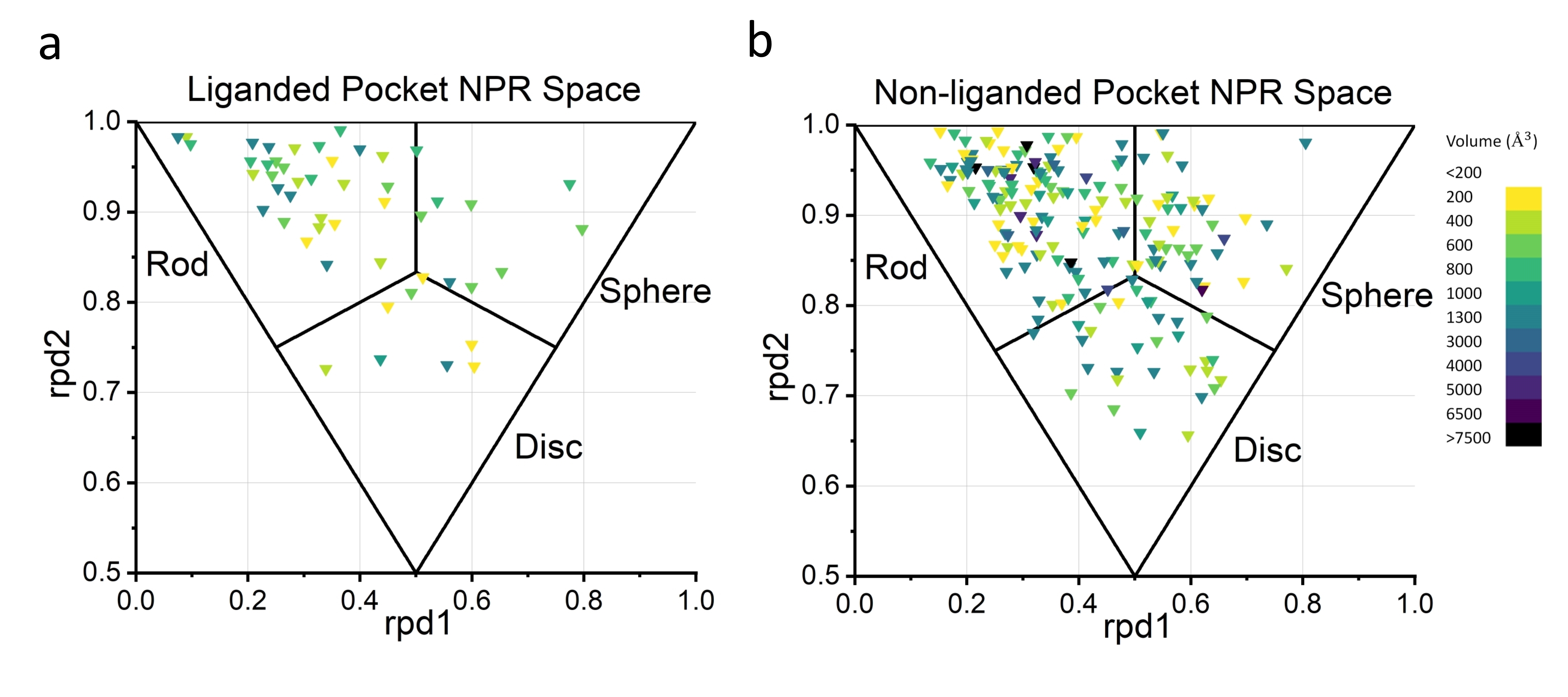
Figure 5. Shape distribution in NPR space of liganded and non-liganded pockets. Color code shows the volume of each pockets.
Some statistical analysis was generated including: the motifs of binding sites, ligands functional groups involved in hydrogen bond interactions and non-bond contacts, the binding motifs of pockets on 2D topology, RNA pocket size distribution and pocket shape distribution. These statistical results could help researchers in the screening of small-molecule drugs.