ZHMolLigGraph Server
ZHMolLigGraph: Exploring Ligand Flexibility in Nucleic Acid Scaffolds Using Graph Neural Networks

About ZHMolLigGraph
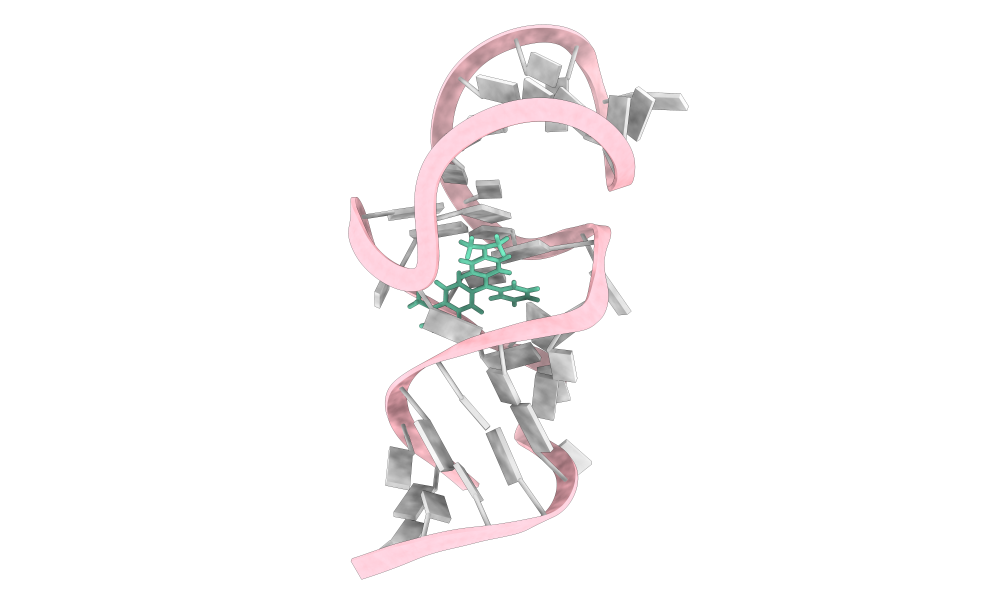
Nucleic acid-ligand interactions are central to gene regulation, disease progression, and therapeutic development. Accurate modeling of these complexes remains difficult because nucleic acid binding pockets are often shallow, irregular, and dynamic, while ligands themselves can undergo substantial conformational changes upon binding. Existing docking approaches usually treat both ligands and nucleic acids as rigid, resulting in sampling pools that lack truly near-native conformations and making it harder for scoring functions to distinguish correct poses from decoys.
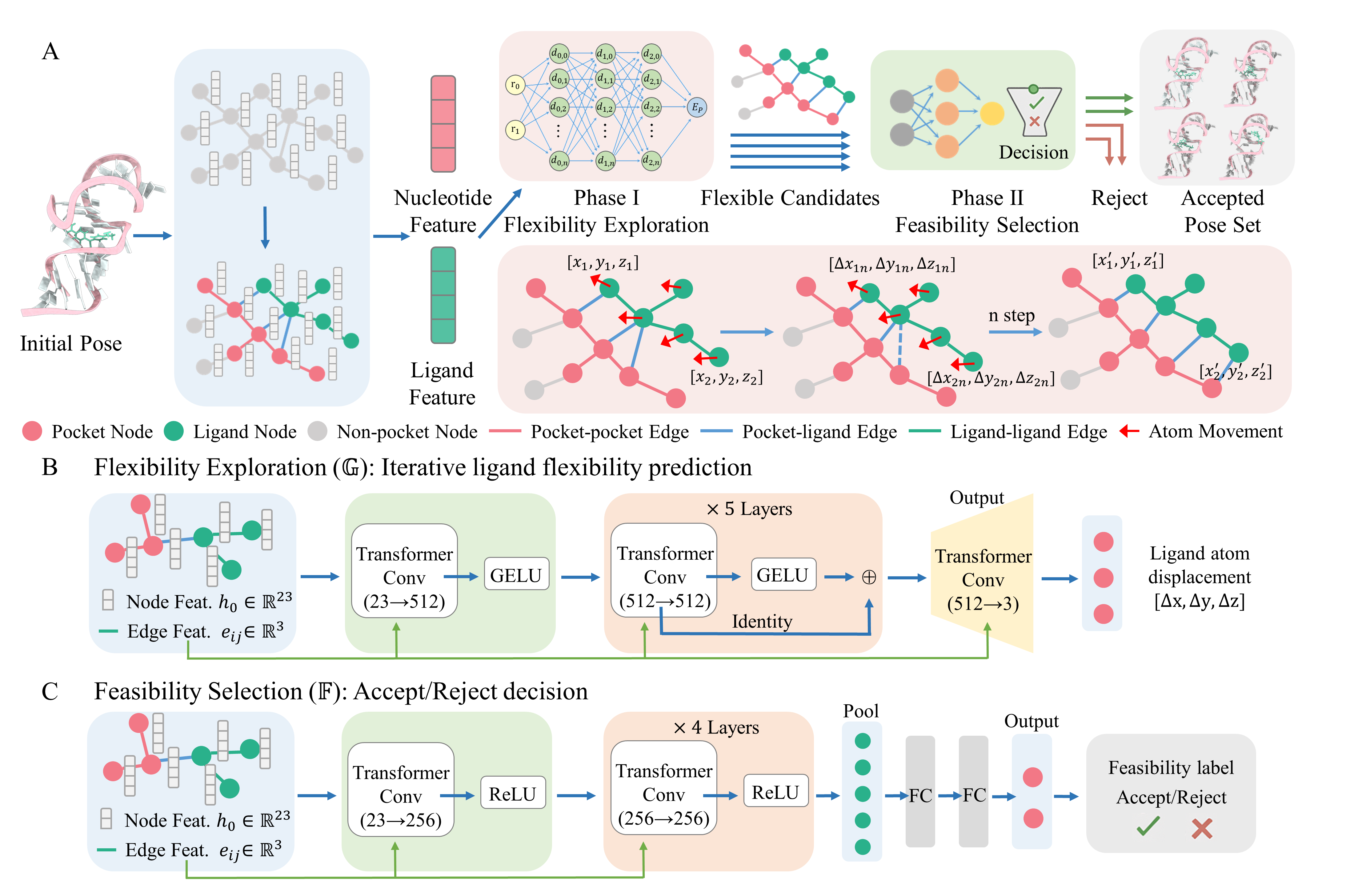
Here we present ZHMolLigGraph, a graph-based deep learning framework that explicitly models ligand flexibility while treating the nucleic acid scaffold as fixed. Each complex is represented as an atomic-level heterogeneous graph without predefined structural features. The framework adopts a two-stage design: Phase I (Flexibility Exploration) applies iterative atomic displacements to explore ligand conformational adaptability, enabling the ligand to traverse a wide range of structural states; Phase II (Feasibility Selection) screens these candidates for geometric and interaction plausibility, retaining only physically reasonable poses rather than ranking all candidates.
Across diverse benchmarks, including local, global, binding site-guided, and conformational-sampling docking, ZHMolLigGraph consistently improved near-native hit rates by 10.30-30.91% compared to conventional docking algorithms, while maintaining fast computational efficiency. By jointly capturing ligand motion and atomic-level interactions against a fixed receptor, ZHMolLigGraph provides a practical and scalable framework for ligand flexibility exploration in nucleic acid-ligand systems. Importantly, the framework is extensible and can be expanded to incorporate receptor flexibility in the future, offering a path toward more faithful modeling of RNA-ligand recognition in realistic biological contexts.
Resources
Related Links
I. 3D Structure Resources
II. RNA Structure Prediction
III. Molecular Docking
IV. Molecular Dynamics
V. Molecular Visualization/Analysis
Visitor Information
| Region | Visit Count | IP |
|---|
Contact
If you have any questions, feel free to contact us at: yjzhaowh@mail.ccnu.edu.cn or cwzengwuhan@mails.ccnu.edu.cn
Institute of Biophysics and Department of Physics, Central China Normal University, Wuhan 430079, China.
References
- Chengwei Zeng, Jiaming Gao, Haoquan Liu, Chen Zhuo, Yunjie Zhao*, Exploring Ligand Flexibility in Nucleic Acid Scaffolds Using Graph Neural Networks. Submitted.